ML-based Diagnosis of Chest Disease
This project was created for Bayer Hacks 2018.
Prizes: 1st Place
Github: Here!
Keynote: Here!
Collaborators: Michael Li
In the 21st century, consumers are bombarded by information at every website they visit. This is useful for companies since it allows opportunities for diverting the customer's attention to browse other products. However, a shift towards consumer-centric UX and app development is needed in order to build a stronger rapport and bring solutions to the customer as fast as possible.
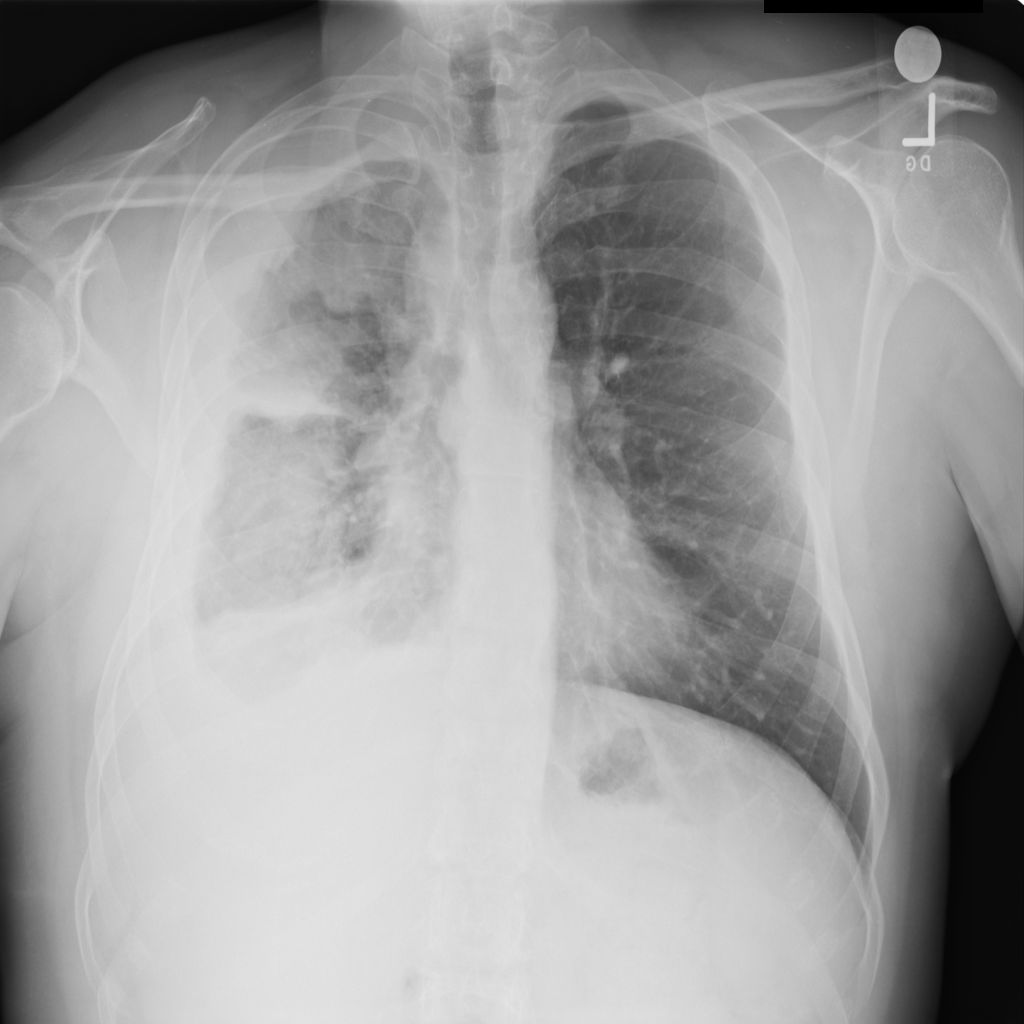
Rxtr is an amalgamation of a powerful 121-level Neural Network and a clean Django front end that allows consumers to classify diseases based on image data inputs. Our solution is extremely focused on the field of Chest Diseases including Pneumonia and Cardiomegaly (14 total). The user simply uploads a chest X-ray and the Pytorch-based backend will produce a probabilistic result based on a training set from the National Institute of Health.
Based on 22-hr training period, the model approaches 80% accuracy. Improvement can be found in increasing convolution/expanding database.
Finally, the application prescribes a medicine from a portfolio for the afflicted disease, creating an end-to-end solution for consumers. Although there is convolution in our Neural Nets, there is no convolution in providing the customer with a solution.
How I Made It:
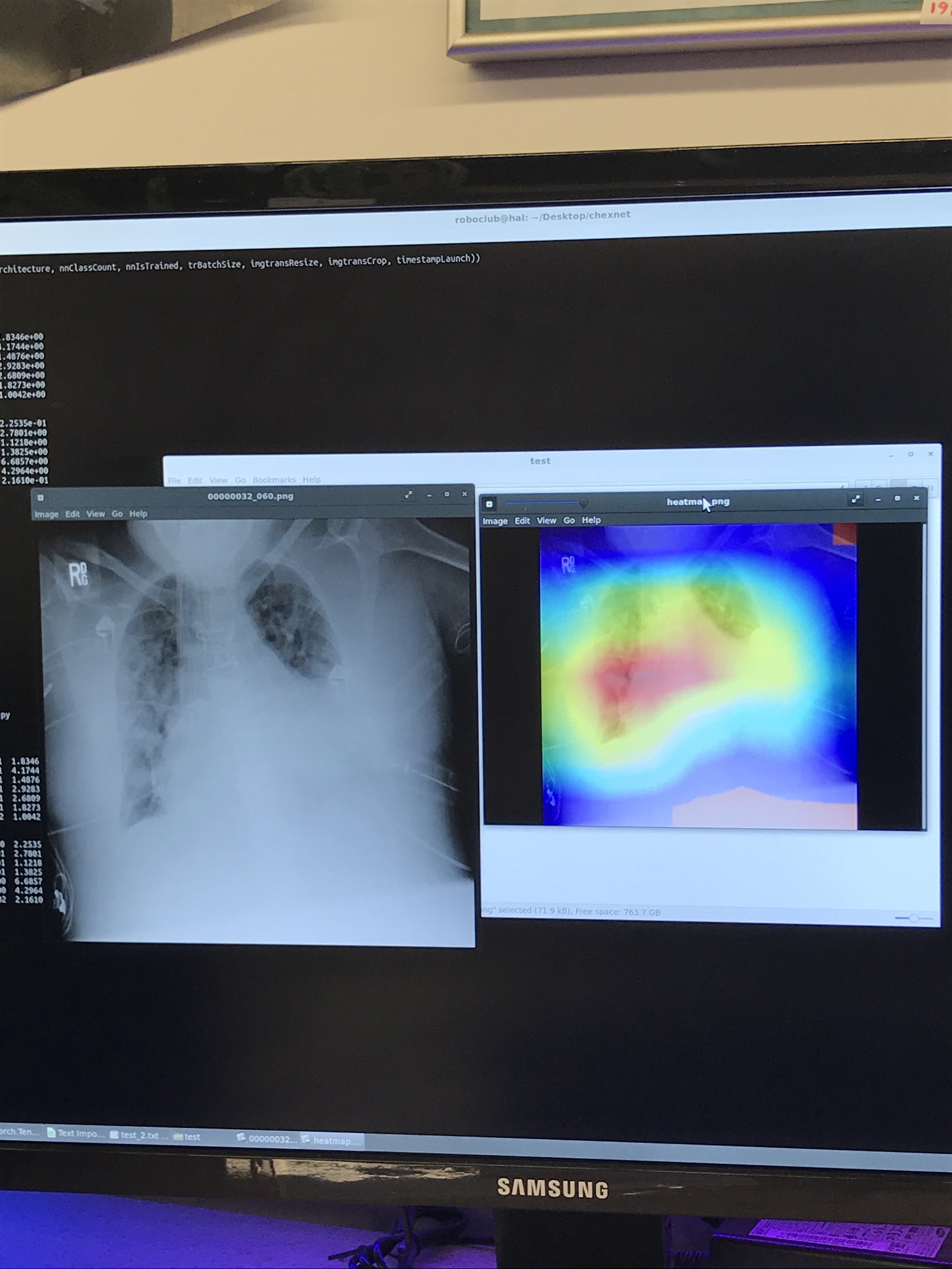
The backend for this application was made using Python. In particular, using a library of machine learning functions and data structure called Pytorch. Pytorch makes it easier to train and manipulate datasets using matrices which are then transformed to a more convenient structure called a tensor.
Furthermore, the model was trained on a computer using an Nvidia Titan V GPU over a 22-hr period. The AUROC (accuracy) score output was then around 80%. Improvements can be seen by using a higher level Neural Network and extending the training period and averaging the outputs.
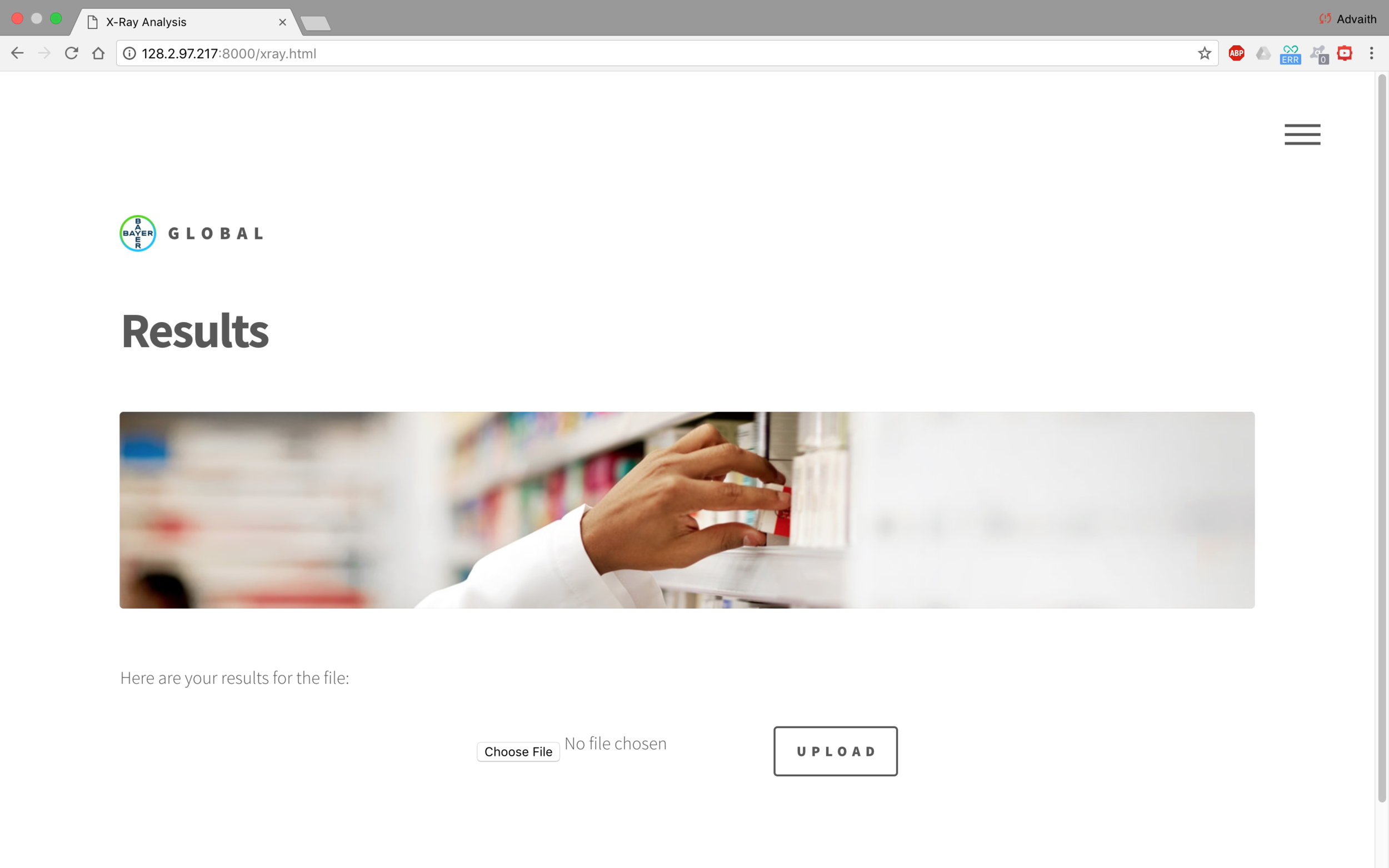
The frontend of the application was developed in Django. It is easy to bootstrap various modules and integrate file upload and hosting. This way, we could blackbox the design process and isolate the various functions of the different modules.
Why I Made It:
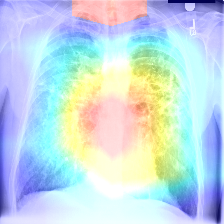
The point of the project was to experiment with various things that the status quo considers to be dominated by humans. Doctors are respected because of the amount of effort they must put to learn and train themselves. The humanoid component of this project is of course the ability to find problematic areas in an image and diagnose a disease based on previously observed images. As a result, the program parallels a human's ability to diagnose and come to a conclusion about a problem.
Furthermore, it is easily integrated into various systems, as you can run the Django server and Python scripts on any server with CUDA capabilities. In the future, I will be experimenting with various other datasets, such as the Cohn-Kanade dataset for emotions, and other medical images.
Furthermore, the system is also easily integrated with something like a chatbot, since you can upload a picture to Facebook and the bot will simply chat you your probable diagnosis. This is more convenient than waiting for a doctor, although the accuracy will need to be worked on. This would complete the humanoid component for the project as it gives the platform an easily accessible user interface.
How it Works:
The image is uploaded to the server, then it is stored in a predefined directory. The path of the image is added to a text file which the Main.py function parses. Then, the image is uploaded to the function, which uses OpenCV to identify various pixel groupings. Then, the outputs from the model are averaged and the diagnosis is written to a text file. The frontend then parses this text file and displays it to the user on the website.
The internals of the machine learning engine are blackboxed but they use principles like Least Squares Approximation and Weight-based Neural Networks to perform the diagnosis.
What Went Wrong:
There were not many hiccups with this project. Most of the time was spent binding the training model to the model that the function calls. However, this was figured out after multiple hours of tinkering with the internal code. Furthermore, multiple modifications had to be made to the Tensor operations in order to not output simply an average score of accuracy but rather the diagnosis based on external images. Multiple OS modifications which included adding and removing paths to the various external modules.
One issue in particular was making the dependencies work with each other. Programming in Python is sometimes a nightmare when using multiple libraries because of cross-version compatibility. The app was made in Python3 but required the OpenCV to run on Python2. As a result, some hacking was required to remove the sys path that hosted OpenCV for Python2 at every iteration of the runtime code.
Dependencies: CUDA, Pytorch, OpenCV, Scikit-Learn, NumPy, SciPy